metabion FAQs
#
metabion international AG
Semmelweisstrasse 3
82152 Planegg
Germany
+49 89 899 363 0
Customer Service
Sales
#
There are two ways of ordering:
- The preferred way is order transmission through our Web Order Portal for most convenient online shopping (please refer here for further details).
- You can order by sending us an e-mail at oligo@metabion.com with our pre-formatted excel order file as attachment. Download respective Order Form.
When you write your e-mail, please make sure to include the following required information in the excel template:
- Name of the oligo/s.
- Sequence of the oligo/s in 5’-3’ orientation. Please indicate any PTO-bond using a star (*).
- Yield range.
- Purification required.
- Delivery form (dry/in water/buffer + concentration).
- Modifications (2'-O-Methyl RNA and 5-Methyl-dC can be selected here).
If you are a new customer, please additionally provide us with:
- Your shipping and billing address.
- Any other information like Purchase Order number, VAT number (VAT only for customers resident in the EU), etc.
In case you choose to transmit orders via e-mail using your own format(s), we need to alert you that above mentioned information in bold print is obligatory for processing your order. Due to extra efforts necessary for individual order format transfer into our system, order processing will take longer as compared to preferred web orders and pre-formatted e-mails. Please note that only files updated to the latest excel/word version (e.g. .xlsx or .docx) are accepted.
How can you convert files to newer formats?
Open the file in a recent Office program and save the file in a newer format. For this go to “File” → “Save as” and choose the newer format from the “Save as type” dropdown menu.
If you want to connect your eProcurement System with our Web Order
Portal (e.g. OCI - Open Catalog Interface), please simply contact our
Customer Service (customerservice@metabion.com).
#
The real answer lies in the resolution limit of the purification method and on the coupling efficiency of the DNA synthesizer. We can synthesize DNA oligos of 220 bases and obtain sufficient quantities by HPLC purification to perform successful gene construction. However, it should be remembered that the longer the oligo, the greater the chance of accumulated sequence errors.
Coupling efficiency is the factor that mainly affects the length of DNA that can be synthesized. Base composition and synthesis scales will also be contributing factors. For more information regarding coupling efficiency, please refer to the FAQ "What is coupling efficiency?".
Importantly, the probability of premature synthesis interruption increases dramatically for longer oligos, due to poor nucleotide-coupling efficiency. Moreover, the synthesis of special “long oligos” requires more raw material, such as amidites and solutions that need to be used in higher excess.
However, as we are very experienced in producing very long oligos, do not hesitate to discuss your projects with our specialists.
#
The current maximum length of standard RNA oligonucleotides is 140 bases. For RNA oligos longer than 140 bases, please inquire.
#
Scale of synthesis refers to the amount of starting Controlled-Pore Glass (CPG) support-bound monomer used to initiate the DNA synthesis, not the amount of final material synthesized. This is the same for all manufacturers of synthetic DNA using standard phosphoramidite chemistry. When a 40 nmole scale synthesis is specified, approximately 40 nmoles of the first base is added to the DNA synthesizer. For an average 25-mer, at least 25% of this starting material will result in failure sequences. Hence it is not possible to produce 40 nmoles of full-length product from a 40 nmole scale synthesis. The losses occur during synthesis, post-synthetic processing, transfer of material, and quality control.
Final yield is the actual amount that we guarantee to deliver.
Please note that OD260 values are a measure of total nucleotides´ optical density. Hence, neither purity nor amount of ordered substance are transparently reflected. For simplification and exemplification reasons look at the following:
1 OD of the 20mer 5´CAT CGT ATT CGA TGC TAC GT 3´
translates into approximately 5 nmol.
1 OD of the 40mer 5´CAT CGT ATT CGA TGC TAC GT CAT CGT ATT CGA TGC TAC GT 3´
translates into approximately 2.5 nmol.
Therefore, a 1 OD guaranteed amount of delivered product can vary significantly, while metabion´s commitment to delivered yields in nmol does not allow for ambiguity in terms of what you expect and pay for.
For oligos of standard length (≤ 33 nucleotides) we guarantee a certain yield of DNA (OD260 applying to a 20-mer) to be delivered according to the respective scale of synthesis and mode of purification.
Our standard custom DNA portfolio features 4 synthesis scales with guaranteed yield expressed in ODs:
* = The yield guarantee is calculated for unmodified oligonucleotides (20mers).
0.02 µmol scale only available for oligos up to 33 bases.
#
Many of the modified amidites are less stable and do not couple as efficiently as the unmodified bases (even though longer coupling procedures may be used). Therefore, failure sequences are more abundant than in normal synthesis. Consequently, most of the modified oligos should be purified by HPLC to remove the more abundant failure sequences. Final yields are reduced as a result of the purification step, which results in a much purer final product, nearly 100% modified.
#
For the 0.2 µmol synthesis scale, monomer coupling is done at a 40-50-fold excess. Doing so for larger scale synthesis (such as for a 1.0 µmol scale) would be cost-prohibitive. Large-scale syntheses are done only at 10-fold mole excess of amidites. To increase the final yields of these large-scale synthesis, the coupling times are extended, so to increase coupling efficiencies.
#
A comprehensive Synthesis Report comes along with each order, indicating oligo name and sequence, composition of bases, modification, base count, purification, yield in ODs, nmol, Tm and molecular weight.
#
This depends on the complexity (length, base composition, modifications) of your requested molecule, as well as on the application desired. Failure sequences may be generated both during and post-synthesis. Due to the nature of synthesis chemistry (coupling efficiency < 100%) and/or post-synthetic modification procedures, there will remain failure sequences (n-x), free modifiers and non-labeled product in the "crude" unpurified product.
We strongly recommend that all modified oligos and oligos longer than 80 nucleotides are subjected to High Pressure Liquid Chromatography (HPLC) purification, no matter which kind of downstream application is envisaged.
For "critical" applications like gene synthesis, subcloning, mutagenesis and alike, HPLC purification should be considered for oligos > 45 nucleotides.
HPLC purification is standard (no additional charge) for all our dual-labeled probes and/or multi-labeled oligos, as well as for all our RNA and large scale oligos.
#
OPC® is an abbreviation for “Oligonucleotide Purification Cartridge”. This purification method consists of a reverse-phase (RP)-chromatography, which separates complete, newly synthesized oligonucleotides from failure sequences, by-products, and other impurities. Specifically, the OPC® cartridge binds the dimithyltrityl (DMT) of the trityl-on oligonucleotide (complete, newly synthesized) from non-DMT bearing molecules. This techniques performs best for oligonucleotides up to 40 nucleotides in length.
Our high pressure liquid chromatography (HPLC) purification system also consists in a RP-chromatography. However, this is performed under high-pressure and highly efficient conditions.
Both in OPC® and HPLC, application of mild acids effectively removes the DMT group, thus allowing elution of purified and detritylated oligonucleotides.
#
Coupling efficiency is a way of measuring how efficiently the DNA synthesizer is adding new bases to the growing DNA chain. If every available base on the DNA chain reacted successfully with the new base, the coupling efficiency would be 100%. Few chemical reactions are 100% efficient. The industry standard for coupling efficiency during DNA synthesis is around 98,5%, with maximum coupling efficiency obtainable being around 99%. This means that at every coupling step approximately at least 1% of the available bases fail to react with the new base being added. Coupling efficiency is significantly influenced by the quality of raw material (amidites and solutions), instruments and synthesis protocols used.
Please note that metabion regularly overperforms the industry standards written above, reaching a coupling efficiency of up to 99,7%, even for long unmodified oligonucleotides.
Moreover, metabion's Quality Control (QC) system ensures that every new batch of chemicals passes strict quality controls. Our machines are serviced by a well organised maintenance program and synthesis cycles are perfectly adjusted to the type of ordered oligo.
#
Coupling efficiency is important because the effects are cumulative during DNA synthesis. The Table below shows the effect of a 1% difference in coupling efficiency and how this influences the amount of full-length product, following the synthesis of oligos of different length. Considering a relatively short oligo of 20 bases, a 1% difference in coupling efficiency can result in a 15% difference, in terms of full-length final product.
The table also shows that the longer an oligo, the lower the yield of full length product that can be expected, due to limitations set by chemistry. Assuming a coupling efficiency of 99% for every single base addition (industry standard is 98.5% in average), the raw product of a 95-mer synthesis would consist of only 38.5% full length oligonucleotide. Separating full length and failure sequences from each other by HPLC purification results in additional loss, so that low yields are a normal matter of fact.
Note that metabion regularly exceeds 99% of coupling efficiency, reaching a coupling efficiency of up to 99,7%, even for long unmodified oligonucleotides.
#
Every DNA base (in terms of DNA synthesis chemistry, we are speaking of phosphoramidite monomers and amidites) added during DNA synthesis has a dimethoxy-trityl (trityl) protecting group attached to the 5´-hydroxyl position. This acid labile trityl-group is bound to the 5’-end of each support-bound monomer and protects the corresponding base from undergoing unwanted chemical reactions during the synthesis cycle. The trityl-group is removed in the first step of each synthesis cycle, immediately before a new base is added, until the elongation of the nucleotide chain is complete. The final trityl-group is removed before delivery (unless otherwise requested).
#
The trityl-group is colorless when attached to a DNA base but it gives a characteristic orange color once removed. The intensity of this color can be measured by UV spectrophotometry and it is directly related to the number of trityl molecules present. Following the first coupling step, the amount of trityl released during deblocking is directly proportional to the amount of full-length oligo synthesized in the previous cycle. When the trityl is cleaved during the deblocking step, the resulting trityl cation is orange in color. The intensity of this color can be measure by UV spectrophotometry. By comparing the intensities of the trityl cation produced after the first and last coupling steps, one can calculate the average successful base coupling per cycle and hence the coupling efficiency.
#
Preparative High Pressure Liquid Chromatography (HPLC) deals with isolating the separated components of a sample, and can be done on small-, mid- and large scale operations. In other words, the objective of a preparative HPLC is isolating and purifying a product. Practically, the sample goes from the detector into a fraction collector or it is collected manually.
Analytical HPLC refers to the processes of separating and identifying the components of a sample. It is usually a small-scale process, whose objective is the qualitative and quantitative determination of a compound. The sample goes from the detector into waste.
metabion offers analytical HPLC as an additional (optional) quality control method, complementing our Mass-Check QC, which is performed by default on all our oligos.
#
metabion is dedicated to reliably deliver high quality products. While every production step is performed in light of achieving best quality, the product is released only if it passes our final inspection. Mass Spectrometry (MS) has become the state-of-the-art technology for verifying the integrity of oligonucleotides, and metabion has been the first custom oligo house who introduced routine mass checks into its operations. Each and every oligo is characterized by either MALDI- or ESI-ToF and stringent release criteria are applied.
Mass Spectrometry allows for the most sensitive detection of low-level by-products/impurities such as:
- n-1/n-x oligos;
- depurination;
- incomplete deprotection;
- acrylonitrile adducts;
- high salt content identification.
Moreover, it is the fastest and most efficient way to identify potential product mix-ups.
We run two different types of Mass Spectrometry instruments in order to cope best with quality and quantity/throughput issues determined by the specifications of the respective oligo/analyte. While each instrument type precisely characterizes oligonucleotides in terms of composition through direct molecular weight measurement, their field of application is diligently adjusted to suitability considerations.
MALDI-ToF instruments typically have a higher throughput, while the limits of using this technique become manifest, if it comes to analyzing long oligonucleotides, or oligos carrying certain photo-labile modifications (e.g.common quenchers like BHQ®s, Dabcyl used in DLPs).
ESI-ToF is less efficient in terms of throughput but perfectly compensates for resolution issues with long oligos as well as for a potential detrimental laser impact on labile/photosensitive modifications – thus being a "natural" complement to MALDI-ToF analysis.
Synthetic oligonucleotide purification is particularly challenging because of the small differences in size, charge and hydrophobicity between the full-length product and impurities, which often co-elute.
For improved analysis of complex samples like long and/or multiple labeled oligos, metabion offers liquid chromatography (LC) coupled with electrospray ionization mass spectrometry (ESI-MS). The mass spectrometer is connected to a high pressure liquid chromatography (HPLC) system, which allows premium analyte characterization via chromatographical separation, followed by respective molecular weight determination. With this system, the mass of oligonucleotides between 2 and 220 bases can be analysed with high accuracy, resolution and sensitivity. Our expert production team will take care of the method (MALDI- or ESI-ToF) that best applies to your sample.
#
Synthesis is monitored through trityl analysis. Quality check by mass spectrometry further ensure top quality. Please visit our FAQs page for more information.
#
For DNA oligos in single tube format
The label on the oligo tube shows basic information like oligo name,
name of person who ordered, oligo sequence including modifications,
oligo ID, amount of DNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more
detailed information on the physical-chemical properties of the oligo,
such as base composition, base count, purification grade, amount of DNA
(OD260 and nmol), Tm and molecular weight. If you have
ordered HPLC purification or your ordered oligo is HPLC purified by
default (e.g. Dual labeled oligos, dual modified oligos, mid- and large scale oligos), you will also get a printout of the preparative chromatogram.
While each and every oligo produced and delivered is characterized by either MALDI- or ESI-ToF before release, Mass-Check documentation/spectra will only be provided if requested at time of order placement. Additional charges may apply. The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either
MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion
internal procedures. This service is free of charge and no printout/PDF
documentation is provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with MALDI-ToF spectra. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with ESI-ToF spectra. Additional charges apply.
Mass Check + Analytical HPLC
Explicitly ordered and performed Mass Check (MALDI-ToF or ESI-ToF
subject to the "nature of the oligo" and metabion internal procedures)
and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For antisense/PTO oligos
The label on the antisense/PTO tube shows basic information like oligo name, name of person who ordered, antisense oligo sequence including modifications, oligo ID, amount of DNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more detailed information on the physical-chemical properties of the oligo, such as base composition, base count, purification grade, amount of DNA in nmol, Tm and molecular weight. Antisense DNA/PTOs are HPLC purified by default and you will also get a printout of the preparative chromatogram.
While each and every oligo produced and delivered is characterized by either MALDI- or ESI-ToF before release, Mass-Check documentation/spectra will only be provided if requested at time of order placement. Additional charges apply.
The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either MALDI- or
ESI-ToF, subject to the "nature of the oligo", and metabion internal
procedures. This service is free of charge and no printout/PDF documentation is
provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with
MALDI-ToF spectra. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with ESI-ToF spectra. Additional charges apply.
For ngs oligos
The label on the oligo tube shows basic information like oligo name, name of person who ordered, oligo sequence including modifications, oligo ID, amount of DNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a technical data sheet containing more detailed information on the physical-chemical properties of the oligo, such as base composition, base count, purification grade, amount of DNA (OD260 and nmol), Tm and molecular weight. You will also get a printout of the Mass-Check documentation/spectrum.
The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion internal procedures. This service is free of charge and no printed/pdf documentation is provided. nGS oligos are always delivered with mass check spectra.
Mass Check + Analytical HPLC
Explicitly ordered and performed Mass Check (MALDI-ToF or ESI-ToF subject to the "nature of the oligo" and metabion internal procedures) and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For DNA/RNA DLPs
The label on the DLP tube shows basic information like oligo name, name of person who ordered, DLP sequence including modifications, oligo ID, amount of DNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more detailed information on the physical-chemical properties of the oligo, such as base composition, base count, purification grade, amount of DNA (OD260 and nmol), Tm and molecular weight. DLPs are HPLC purified by default and you will also get a printout of the preparative chromatogram. Additionally, you will receive a hard copy of the mass check spectrum.
The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either
MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion
internal procedures. This service is free of charge and you will receive
a hard copy of the Mass Check spectrum.
Mass Check + Analytical HPLC
Explicitly ordered and performed Mass Check (MALDI-ToF or ESI-ToF
subject to the "nature of the oligo" and metabion internal procedures)
and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For DQP/LNA/MGB probes
A hardcopy of the preparative HPLC chromatogram and the mass spectrum are included in addition to the respective synthesis report and delivery note.
For LC® probes
The label on the probe tube shows basic information like oligo name, name of person who ordered, probe sequence including modifications, probe ID, amount of DNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more detailed information on the physical-chemical properties of the oligo, such as base composition, base count, purification grade, amount of DNA (OD260 and nmol), Tm and molecular weight. LC®probes are always HPLC purified and you will also get a printout of the preparative chromatogram.
The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either
MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion
internal procedures. This service is free of charge and you will receive
a hard copy of the Mass Check spectrum.
Mass Check + Analytical HPLC
Explicitly ordered and performed Mass Check (MALDI-ToF or ESI-ToF
subject to the "nature of the oligo" and metabion internal procedures)
and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For ZNA primers and probes
On the tube label you will find basic information such as oligo name, name of the person who placed the order, oligo sequence including modifications, oligo ID, delivered quantity of DNA and molecular weight.
In addition, you will receive a technical data sheet including more detailed information on physico-chemical properties of the oligo like base composition, GC-content, synthesis scale, purification grade, quality control information, etc. You will also receive a printout of the Mass-Check, free of charge.
For dsDNA/duplex DNA
The label on the dsDNA/duplex DNA tube shows basic information like oligo name, name of person who ordered, DLP sequence including modifications, oligo ID, amount of DNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more detailed information on the physical-chemical properties of the ssDNA oligo, such as base composition, base count, purification grade, amount of DNA (OD260 and nmol), Tm and molecular weight. dsDNA/duplex DNAs are HPLC purified by default and you will also get a printout of the preparative chromatogram.
While each and every oligo produced and delivered is characterized by either MALDI- or ESI-ToF before release. Mass-Check documentation/spectra will only be provided if requested at time of order placement. Additional charges may apply.
For RNA oligos
The label on the oligo tube shows basic information like oligo name, name of person who ordered, oligo sequence including modifications, oligo ID, amount of RNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more
detailed information on the physical-chemical properties of the oligo,
such as base composition, base count, purification grade, amount of RNA
(OD260 and nmol), Tm and molecular weight.
Each and every
oligo produced and delivered is analyzed by either MALDI- or ESI-ToF
before release, and Mass-Check documentation/spectra will be provided
upon additional charges (desalted RNA) or free of charge (HPLC purified
RNA). The following terminology is used for differentiating between
offered QC:
For desalted RNA:
Mass Check
Standard quality control performed on
each and every oligo. Either MALDI- or ESI-ToF, subject to the "nature
of the oligo", and metabion internal procedures. This service is free of
charge and no printed/pdf documentation is provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with MALDI-ToF spectra. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with ESI-ToF spectra. Additional charges apply.
Analytical HPLC
Performed Mass Check and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC but without MS spectra. Additional charges apply.
Mass Check + Analytical HPLC
Explicitly ordered
and performed Mass Check (MALDI-ToF or ESI-ToF subject to the "nature of
the oligo" and metabion internal procedures) and Analytical HPLC (see
FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For HPLC purified RNA:
Mass Check
Standard quality control performed on
each and every oligo. Either MALDI- or ESI-ToF, subject to the "nature
of the oligo", and metabion internal procedures. This service is free of
charge and will always be delivered with hard copies of the Mass Check
spectrum for all our HPLC purified RNA products.
Analytical HPLC
Mass Check (MALDI-ToF or ESI-ToF
subject to the "nature of the oligo" and metabion internal procedures)
and explicitly ordered Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For RNA Longmers
The label on the oligo tube shows basic information like oligo name, name of person who ordered, oligo sequence including modifications, oligo ID, amount of RNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a synthesis report containing more
detailed information on the physical-chemical properties of the oligo,
such as base composition, base count, purification grade, amount of RNA
(OD260 and nmol), Tm and molecular weight.
Each and every oligo produced and delivered is analyzed by either MALDI-
or ESI-ToF before release, and Mass-Check documentation/spectra will be
provided upon additional charges (desalted RNA) or free of charge (HPLC
purified RNA). The following terminology is used for differentiating
between offered QC:
For desalted RNA:
Mass Check
Standard quality control performed on each and every oligo. Either
MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion
internal procedures. This service is free of charge and no printed/pdf
documentation is provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with a MALDI-ToF spectrum. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with an ESI-ToF spectrum. Additional charges apply.
Analytical HPLC
Performed Mass Check and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC but without MS spectra. Additional charges apply.
Mass Check + Analytical HPLC
Explicitly ordered and performed Mass Check (MALDI-ToF or ESI-ToF
subject to the "nature of the oligo" and metabion internal procedures)
and Analytical HPLC (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For HPLC purified RNA:
Mass Check
Standard quality control performed on each and every oligo. Either
MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion
internal procedures. This service is free of charge and will always be
delivered with hard copies of the Mass Check spectrum for all our HPLC
purified RNA products.
Analytical HPLC
Mass Check (MALDI-ToF or ESI-ToF subject to the "nature of the oligo"
and metabion internal procedures) and explicitly ordered Analytical HPLC
(please read FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
For siRNA/RNAi/duplex RNA
The label on the siRNA tube shows basic information like oligo name, name of person who placed the order, siRNA sequence including modifications, oligo ID, amount of RNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a technical data sheet containing more detailed information on the physical-chemical properties of the oligo, such as base composition, base count, purification grade, amount of RNA in nmol, Tm and molecular weight. Additionally, you will also receive a hard copy of the Mass-Check documentation/spectrum of each single strand of the duplex. The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either MALDI- or
ESI-ToF, subject to the "nature of the oligo", and metabion internal
procedures. This service is free of charge and the Mass Check spectrum will be
provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with
MALDI-ToF spectra. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with ESI-ToF spectra. Additional charges apply.
For MTP oligos
The label on the oligo plate shows basic information like name of person who ordered, name of the plate and order ID.
In addition, you will receive a synthesis report containing more detailed information, such as name of the oligos, physical-chemical properties of the oligos, such base composition, base count, purification grade, amount of DNA (OD260 and nmol), Tm and molecular weight.
While each and every oligo produced and delivered is characterized by either MALDI- or ESI-ToF before release, Mass-Check documentation/spectra will only be provided if requested at the time of order placement. Additional charges may apply. The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either
MALDI- or ESI-ToF, subject to the "nature of the oligo", and metabion
internal procedures. This service is free of charge and no printed/pdf
documentation is provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with MALDI-ToF spectra. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with ESI-ToF spectra. Additional charges apply.
Mass Check + Analytical HPLC
Explicitly ordered and performed Mass Check (MALDI-ToF or ESI-ToF
subject to the "nature of the oligo" and metabion internal procedures)
and Analytical High Pressure Liquid Chromatography (see FAQ "What is the difference between preparative and analytical HPLC?"). Product delivered with analytical HPLC and MS spectra. Additional charges apply.
#
Our default delivery form is dry. This ensures maximum stability during transport and storage.
#
We recommend to dissolve single stranded RNAs in 1x TE buffer (10 mM TrisCl, pH7.5, 0.1 mM EDTA; prepared under RNase-free conditions). This buffers the pH and chelates metal ions, which may contribute to RNA degradation. RNase-free water is also acceptable.
metabion's RNA oligonucleotides are delivered deprotected and purified – ready to use. They are RNAse free, but as RNA is highly susceptible to degradation by exogenous RNAses introduced during handling, it is essential that you conduct all handling steps under sterile, RNAse free conditions. Never handle RNA without wearing gloves. RNAse free reagents, barrier pipette tips and tubes should be used!
For more information, please read our guidelines.
#
Please have a look to our guidelines.
#
To gain a maximum shelf life for oligonucleotides, samples should generally be stored dehydrated at ≤ –15 °C in absence of light. Under the mentioned conditions, samples are stable for at least 6 months. In case of a longer storage period, oligos should be pretested for molecular integrity prior to experimental use. If stored frozen at –20 °C or –70 °C, it will remain stable for several months. Repeated freeze-thaw should be avoided, as this will denature the oligo. Moreover, the oligo stability in solution depends on the pH. Dissolving oligos into acidic solutions may result in oligo degradation. Therefore, avoid the use of distilled water, since solution pH may be as low as 4-5.
In addition to what above advised, we recommend that you
minimize the exposure of modified oligonucleotides– especially those
fluorescently labelled - to light, to avoid any bleaching effect.
Moreover, we recommend storing dye-labelled oligos highly concentrated and not
in working dilutions, if you are not planning to use them within 24 hours. The
higher the dilution factor, the faster the fluorescent activity fades away.
Therefore, try to store highly concentrated aliquots frozen, thaw them only
once, dilute them just before use and store the aliquots at 4 °C in the dark.
#
The best way to store RNA is as a dry pellet at -20 °C to -80 °C and to avoid frequent freeze-thaw cycles. If you want to store your RNA in solution re-suspend the delivered pellet in an RNAse-free solution buffered at pH 7.4 - 7.6 and store at -20 °C or less. We recommend that RNAs are re-suspended at a convenient stock concentration and stored in small aliquots to avoid multiple freeze thaw cycles.
For modified oligonucleotides – especially for fluorescent dye labelled oligos – you should minimize the probe’s exposure to light because of its bleaching effect. Additionally, we recommend storing dye labelled oligos highly concentrated, rather than in working dilutions, if you don’t use them immediately. The higher the dilution factor the faster the fluorescent activity decreases/fluorescence fades away. Therefore, try to store highly concentrated aliquots frozen, thaw them only once, and dilute them just before usage. Click here to download our guidelines for handling RNAs.
#
We recommend that each aliquot of RNA should not be stressed by more than 4 - 5 freeze-thaw cycles to ensure product integrity.
#
To gain a maximum shelf life for oligonucleotides, samples should generally be stored dehydrated at ≤ -15 °C in absence of light. Under the mentioned conditions, samples are stable for at least 6 months. In case of a longer storage period, oligos should be pretested for molecular integrity prior to experimental use. If stored frozen at –20 °C or –70 °C, it will remain stable for several months. Repeated freeze-thaw should be avoided, as this will denature the dsDNA. Moreover, the dsDNA stability in solution depends on the pH. Dissolving dsDNA into acidic solutions may result in its degradation. Therefore, avoid the use of unpurified distilled water as a diluent, since solution pH may be as low as 4-5. In addition to what above advised, we recommend that you minimize the exposure of modified dsDNA/duplex DNA – especially if fluorescently labelled - to light, to avoid any bleaching effect.
Moreover, we recommend storing dye-labelled dsDNA/duplex DNA highly concentrated and not in working dilutions, if you are not planning to use it within 24 hours. The higher the dilution factor, the faster the fluorescent activity fades away. Therefore, try to store highly concentrated aliquots frozen, thaw them only once, dilute them just before you use the probe and store the aliquots at 4 °C in the dark.
#
We recommend to store ZNA®s at –20 °C in TE buffer (pH 7.4). For modified – especially fluorescently labelled-oligonucleotides – please avoid or minimize exposure to light, to avoid any bleaching effect. Also we recommend storing dye-labelled oligos highly concentrated, unless you use the working dilutions within 24 hours. The higher the dilution factor, the faster the fluorescent activity fades away. Therefore, keep highly concentrated aliquots in the freezer, thaw them preferably only once, dilute them just before use. Store the thawn aliquot in the dark. Under these conditions, the ZNA oligos will remain stable for several months. Repeated freeze-thaw cycles must be avoided, as this will denature the oligo and compromise the integrity of any added label. Avoid the use of non-sterile distilled water, since its pH may be as low as 4–5. Heating ZNA® in basic buffers induces spermine cleavage and must be avoided.
#
To gain a maximum shelf life for oligonucleotides, samples should generally be stored dehydrated at ≤ –15 °C in absence of light. Under the mentioned conditions, samples are stable for at least 6 months. In case of a longer storage period, oligos should be pretested for molecular integrity prior to experimental use. If a sterile solution (e.g. water, biological buffer) is used as diluent, the re-suspended the probe will be stable at 20 °C for several days to weeks, at 4 °C for about a month. If stored frozen at –20 °C or –70 °C, it will remain stable for several months.
For correct storing and best performance of your probe, we recommend the following:
- Avoid repeated freeze-thaw, as this will denature the probe;
- Avoid the use of distilled water as a diluent, since its pH may be as low as 4–5. The probe stability in solution depends on the pH. Dissolving probes into acidic solutions may result in oligo degradation. Therefore, use purified distilled water;
- Minimize the exposure of fluorescent probes to light, to avoid any bleaching effect;
- Store probes highly concentrated and not in working dilutions, if you are not planning to use them within 24 hours. The higher the dilution factor, the faster the fluorescent activity fades away. Therefore, try to store highly concentrated aliquots frozen, thaw them only once, dilute them just before you use the probe and store the aliquots at 4 °C in the dark.
#
DNA oligos in single tube format
Unmodified standard oligos (≤33 bases) ordered before 6 pm German time will be shipped the next working day. HPLC purified, longer and modified oligos need an additional working day to be ready for shipping.
Shipping time within Europe does normally not exceed 1 day. For countries outside of Europe, please inquire.
RNA oligos, the expected turnover time is:
- desalted RNA oligonucleotide in single tube format is 3–5 days;
- RNA in plates, expected turnover time is 5–7 business days*;
- HPLC-purified, unmodified or single-labelled RNA oligonucleotides from our standard portfolio are usually processed within 5–10 business days including purification and quality control*;
- duplex oligos take about 10–15 days to be produced, purified and QCed*;
- for RNA dlps you should consider 10–15 days as a realistic turnaround time and for RNA longmers depending on sequence length/complexity and modification level 8-15 business days*.
*Freight forwarders delivery time to the final destination is not included in all cases. In terms of "counting" working days, orders placed past 15:00 (Munich time) are considered to be next day's order. Be assured that we try to process your order as quickly as possible without compromising on quality!
The expected average in-house turnover time for antisense/PTOs is 3–4 working days.
For DLPs, LC®probes, MGB probes: the expected average in-house turnover time is 4–5 working days. For RNA DLPs, the expected average in-house turnover time is 10–15 working days.
For dsDNA oligos and siRNAs: the expected average in-house turnover time is 5–10 working days.
RNA longmers are usually processed within 8–15 business days including purification and quality control, depending on sequence length/complexity and modification level.
For ngs oligos: standard in-house turnover time is 2–4 working days.
Typical in-house turnover time for DNA Oligos in MTP format are 2-4 working days. HPLC purified, long and modified oligos need an additional working day to be ready for shipping.
Please note that we perform strict quality controls on your ordered oligos. In case one or more oligos do not pass our quality control, they will have to be resynthesized. This may, of course, result in a delay.
#
Of course we can and we would be happy to do so! Please inquire for feasibility and a quotation; write us an email at info@metabion.com.
#
Oligos are made using a DNA synthesizer, which is basically a computer-controlled reagent delivery system. The first base is attached to a solid support, usually a glass or polystyrene bead, which is designed to anchor the growing DNA chain in the reaction column. DNA synthesis consists of a series of chemical reactions.
Each cycle of reactions results in the addition of a single DNA base. A chain of DNA bases can be built by repeating the synthesis cycles until the desired length is achieved.
#
The chemistry of RNA synthesis is identical to the DNA synthesis except for the presence of an additional protecting group at the 2' hydroxyl position of ribose. This position is protected with silyl groups (usually TBDMS), which are stable throughout the synthesis. The remaining positions on both the sugar and the bases are protected in the same fashion as in DNA. Please visit our FAQs for more information.
#
Please note, that for custom made products an order/item cancellation or modification free of charge is only granted within a grace period of 30 minutes from the air date of this confirmation in order to guarantee fast delivery times.
For "off-the-shelf" products, cancellation/modification of orders/items is also accepted within a grace period of 30 minutes from the air date of the respective order confirmation.
Later cancellations/modifications are principally non-refundable, however a partial refund might be possible depending on the status of order processing. We will inform you accordingly.
Any cancellation/modification of an order needs to be notified in writing. If you've ordered via our Web Ordering Portal, you can cancel by visiting your order history.
If you've ordered via e-mail, you can send us an e-mail to cancellationnotification@mymetabion.com with the subject "cancel N230502-079".
Please send any other questions or requests to our customerservice at customerservice@mymetabion.com.
#
While some manufacturers state that re-annealing of complementary DNA oligonucleotides is generally not necessary, we recommend to go through the process of duplex formation after having dissolved the delivered ds oligonucleotide prior to your experiments applying a suitable protocol.
For your convenience, we recommend to follow our guidelines prepared for download.
#
While some manufacturers state that re-annealing is generally not necessary, we do recommend to re-anneal the dissolved duplex using a suitable protocol. For your convenience, we recommend to follow our guidelines prepared for download.
#
MGB (Minor Groove Binder), ZNA® and LNA (Locked Nucleic Acids) are known to increase the Tm of an oligo sequence.
MGB probes include a minor groove binder moiety at the 3’ end that increases the melting temperature (Tm) of the probe and stabilizes the hybridization of the probe DNA to its target sequence. The introduction of the minor groove binder moiety is sequence-independent, meaning the core sequence remains “unmodified”.
ZNA®s are oligonucleotides conjugated with repeated cationic spermine units that decrease electrostatic repulsions with target nucleic acid strands, and greatly improve hybridization properties by enhanced affinity to the complementary target sequence as well as increased stability of the formed duplex at an unprecedented specificity. The “Tm boost” generated by adding ZNA® to either end of the oligonucleotide probe is significant and also sequence-independent.
In contrast LNA probes rely on modified nucleotide chemistry with regard to the sugar component involved, while the organic base component is “unmodified” and thus follows Watson-Crick base-pairing rules when mixed with DNA or RNA bases in an oligonucleotide. When incorporated into an oligonucleotide probe, locked nucleic acid monomers increase structural stability, resulting in a raise of the formed duplex´melting temperature (Tm). Locked Nucleic acids are not recognized by DNA/RNAses as a substrate, hence LNA modified oligonucleotides also display significant resistance to nucleases.
In summary, metabion offers all three duplex stability enhancing modifications, and therefore provides greatest flexibility in assay design and choosing the right option for your required application.
#
MGB (Minor Groove Binder) and LNA (Locked nucleic acids) bases are known to alter the Tm of a oligo sequence. ZNA® shows similar properties concerning increased affinity to their target (Tm increase), especially when it comes to short sequence probes. Additionally, ZNA® has unique features like the enhanced/accelerated target recognition, which can be used to shorten cycling times or for improved quantification accuracy of low abundant transcripts.
You will find approximations about the Tm increasing effect of MGB and LNA, but an exact calculation of the final Tm is not possible, and cannot be provided a priori. The effect of Tm increase is highly dependent on sequence composition and GC content; therefore, each assay has to be optimized.
MGB is often supported by “Superbases”, which increase affinity.
The same applies to ZNA, and metabion offers Tm modifying bases to complement the use of ZNA primers and probes:
- C-5 propynyl-dC (pdC) raising melting temperature by ~2.8 °C per substitution
- C-5 propynyl-dU (pdU) raising melting temperature by ~1.7 °C per substitution.
In summary, ZNA probes provide broad flexibility in assay design and represent therefore an effective alternative to MGB- and LNA -containing oligonucleotides.
Of course ZNA® will not be the solution for all your needs and applications, nor do MGB or LNA. It is a wonderful and promising technology, which has to be explored. Be a part of it!
#
Coupling efficiency during solid phase synthesis is the major factor that limits length and/or quality of the stepwise built molecule. Moreover, any modifications/labels added to or incorporated into the nucleotide chain affect the integrity of the oligomer. Sequence composition and synthesis scales are contributing factors as well. We routinely offer ZNA®s of 40 nucleotides in length. Minimum standard length is 8 nucleotides (for probes), and 10 nucleotides (for primers).
If your needs are outside this range, don’t hesitate to discuss your projects with our specialists.
#
Using our optimized production pipeline, we can deliver probes of over 40 bases. Please inquire.
#
Using our optimized production pipeline, we can deliver MGB probes of over 40 bases. The maximum length of an oligonucleotide depends both on the sequence and on the modifications of interest. For example, consider a probe having a reporter at the 5’-end and a quencher at the 3’-end. The length of the probe can compromise the activity of the quencher, if the distance between reporter and quencher is too high. Therefore, we consider a length of 40 bases as maximum, for an MGB probe to function optimally.
However, you can now go beyond this limit with our ZNA technology! Want to learn more? Click here.
With this technology, the quencher is brought in closer proximity to the dye and its effect is optimized. You only need to take into account that the Tm of the resulting probe will be higher, compared to that of a native (non-ZNA) probe. To get best performance for your experiments, we can help you designing your probe. Please do not hesitate to contact us!
#
Using our optimized production pipeline, we can deliver DLPs of over 40 bases. The maximum length of an oligonucleotide depends both on the sequence and on the modifications of interest. For example, consider a DLP having a reporter at the 5’-end and a quencher at the 3’-end. The length of the DLP can compromise the activity of the quencher, if the distance between reporter and quencher is too high. Therefore, we consider a length of 40 bases as maximum, for a DLP probe to function optimally.
However, you can now go beyond this limit with our ZNA technology! Want to learn more? Click here.
With this technology, the quencher is brought in closer proximity to the dye and its effect is optimized. You only need to take into account that the Tm of the resulting probe will be higher, compared to that of a native (non-ZNA) probe. To get best performance for your experiments, we can help you designing your probe. Please do not hesitate to contact us!
#
5 yield ranges based on length independent delivered quantities* for probes from 18-40 nucleotides:
- ≥ 1 < 5 nmol
- ≥ 5 < 10 nmol
- ≥ 10 < 20 nmol
- ≥ 20 < 30 nmol
- ≥ 30 < 50 nmol
- ≥ 50 nmol
* Please note that the term “yield range” refers to the final amount of product you will actually receive. Instead, OD260 values are a measure of total nucleotides' optical density. Hence, neither purity nor amount of ordered substance are transparently reflected. For simplification and exemplification reasons look at the following:
1 OD of the 20mer 5´cau cgu auu cga ugc uac gu 3´ translates into approximately 5 nmol;
1 OD of the 40mer 5´cau cgu auu cga ugc uac guc auc gua uuc gau gcu acg u 3´ translates into approximately 2.5 nmol.
#
For RNA oligonucleotides between 5-40 nts and for a quantity delivered between 10-100 nmol, we offer RNA oligonucleotides desalted as well as HPLC purified. Over 100 nmol and 40 nts our default purity is HPLC! Such RNA require HPLC purification to guarantee the top quality required for your experiments. For such RNA oligonucleotides, HPLC purification is included in our price per base/probe/duplex. Visit our Standard Portfolio.
#
Given the use of siRNA oligos in cells and in vivo, high purity is essential.
High pressure liquid chromatography (HPLC) purification is standard (no additional charge) for siRNAs. For applications in cells and in vivo, we recommend an additional Size Exclusion Chromatography (SEC) purification. This is a purification that removes salts and other small molecules, whose quantity and/or quality can be toxic for cells.
#
For RNA oligonucleotides between 5–40 nts and for a quantity delivered between 10–100 nmol, we offer RNA oligonucleotides desalted as well as HPLC purified. Over 100 nmol and 40 nts our default purificatiom method is HPLC! Such RNA require HPLC purification to guarantee the top quality required for your experiments. For such RNA Longmers, HPLC purification is included in our price per base/probe/duplex. Visit our Portfolio.
#
This depends on the complexity (length, base composition, modifications) of your requested molecule, as well as on the application desired. Failure sequences may be generated both during and post-synthesis. Due to the nature of synthesis chemistry (coupling efficiency < 100%) and/or post-synthetic modification procedures, there will remain failure sequences (n-x), free modifiers and non-labeled products, respectively, in the "crude" unpurified product.
High pressure liquid chromatography (HPLC) purification is standard (no additional charge) for dsDNA/duplex DNA. For applications in cells and in vivo, we recommend an additional Size-Exclusion Chromatography (SEC). This is a size exclusion chromatography purification that removes salts and other small molecules, whose quantity and/ or quality can be toxic for cells (please inquire).
#
Given the use of antisense oligos in cells and in vivo, high purity is essential.
High pressure liquid chromatography (HPLC) purification is standard (no additional charge) for antisense/PTOs. For applications in cells and in vivo, we recommend an additional Size-Exclusion Chromatography (SEC) purification. This is a size exclusion chromatography purification that removes salts and other small molecules, whose quantity and/or quality can be toxic for cells.
#
All ZNA® oligos are routinely analyzed by checking and verifying the molecular weight by Mass Check analysis (ESI- or MALDI-ToF) for actual molecular weight determination and comparing the result with to the theoretical and to be expected mass. If practice meets theory applying our strict quality criteria are the basis for final product release.
Mass Check analysis (ESI- or MALDI-ToF) is routinely performed on all ZNA®s. If the theoretical expected mass matches the molecular weight determined by the mass check, the product is released.
#
Purified distilled water, TE or any biological buffers (i.e. with physiological pH) are acceptable as diluents. The recommended diluent volume is 100 µl - 1 ml, the concentration depending on the application to be used and the yield of the resulting product (see also FAQ "How stable is my probe once I have resuspended it?").
Standard concentration for PCR primers is 0.1 mM.
#
DNA synthesis is a complicated process, which has improved significantly over the last years. Despite these improvements, all manufacturers have an inherent failure rate. We are constantly developing our processes and systems to minimize these losses; however, it is inevitable that we will occasionally have to re-synthesize some oligos. Please note that metabion performs strict quality controls on each and every oligo synthesized. If an oligo does not pass our quality tests, it will be resynthesized.
#
There is a normal degree of variation in the appearance of the supplied dry oligonucleotide pellets. Variation in appearance per se does not indicate a quality defect. In general, appearance of unmodified and dye-labeled oligo pellets may vary from powdery to hyaloid. The color of unmodified oligo pellets may range from transparent over off-white and yellowish to tan. The pellets of labeled oligos are colored according to the dye attached.
#
Unless requested, oligos are synthesized without either 3´or 5´ phosphate. The 5´ and/or 3’-phosphate is available as a modification at additional charge.
#
Unless requested, dsDNA are synthesized with neither 3´nor 5´ phosphate. Both, the 5´and the 3´end will carry a Hydroxyl-group. The 5´ and/or 3’-phosphate is available as a modification at additional charge.
#
Unless requested, siRNA are synthesized with neither 3´nor 5´ phosphate. Both, the 5´and the 3´-end will carry a Hydroxyl-group. The 5´ phosphorylation of both strands is available as a modification at additional charge.
#
Unless requested, antisense/PTO are synthesized with neither 3´nor 5´ phosphate. Both, the 5´and the 3´-end will carry a Hydroxyl-group. The 5´ and/or 3’-phosphate is available as a modification at additional charge (please inquire).
#
Unless requested, oligos are synthesized without either 3´or 5´ phosphate. The same goes for ZNA® oligos. The 5´ and/or 3’-phosphate is available as a modification at additional charge.
#
Ligation reactions require a 5´phosphate. If your oligos do not contain a 5´ phosphate, ligation will not occur or only with a very low efficiency. The problem can be addressed without ordering an additional oligo pair: phosphorylate your oligos enzymatically with kinase before use in ligation reactions.
#
First of all, we strongly recommend our customers to sequence more than one clone and compare the sequencing results. In most cases, sequencing at least 3 clones is sufficient to find the clone with the desired sequence. This is also due to circumstances out of oligo manufacturers´ control, like sequence errors generated by enzymes like Taq polymerase used in downstream experiments. PCR-cloned sequences may contain errors due to the inherent "infidelity" of any kind of available polymerases. Taq may have error rates as high as 0.25%. If Taq was not used, the difference could be due to a recombinant vector or the host cell system "self-correcting" errors.
The problem can also be due to the primers. Base insertions are attributed to a small amount of detritylated amidite present during coupling, while deletions are probably due to failure sequences that did not get capped and were subsequently extended.
However, a better explanation for the observation of altered sequences is the incomplete deprotection of the oligo. If an oligo still bears a protecting group in one or more positions, this will be transferred to the subsequent PCR product, which is then transformed into E. coli. Here, the host mismatch repair system will likely attempt to correct the corresponding anomaly with a base, which might be the wrong one. The most likely culprit for incomplete deprotection is the isobutyryl protected dGs. These are the most difficult protection groups to remove.
In general, the longer the oligo, the greater the probability of side reactions during oligo synthesis, along with higher chances to incur into incomplete deprotection. Potential sources of side reactions causing failure products are depurination (which mainly affects the base A) and formation of secondary structures due to the oligos’ sequence. There is no way to completely exclude these effects! However, metabion tries to minimise these failures by continuously optimising synthesis as well as purification protocols!
#
Base insertions are attributed to a small amount of detritylated
amidite present during coupling, while deletions are probably due to failure
sequences that did not get capped and were subsequently extended.
However, a better explanation for the observation of altered sequences is the
incomplete deprotection of the oligo. If an oligo still bears a protecting
group in one or more positions, this will be transferred to the subsequent PCR
product, which is then transformed into E.coli. Here, the host mismatch repair
system will likely attempt to correct the corresponding anomaly with a base,
which might be the wrong one. The most likely culprit for incomplete
deprotection is the isobutyryl protected dGs. These are the most difficult
protection groups to remove.
In general, the longer the oligo, the greater the probability of side reactions
during oligo synthesis, along with higher chances to incur into incomplete
deprotection. Potential sources of side reactions causing failure products are
depurination (which mainly affects the base A) and formation of secondary
structure due to the oligos’ sequence. There is no way to completely exclude
these effects! However, metabion tries to minimise these failures by
continuously optimising synthesis as well as purification protocols!
#
Yes, they are as follows:
#
You may want to consider the following:
#
For a detail explanation of DLPs and their use in qPCR, or for more information regarding how to design probes for qPCR, please refer to our Knowledge-hub.
You also need to consider the following:
#
Yes, they are as follows:
#
For a detail explanation of LC® probe and their use in qPCR, please refer to our Knowledge-hub.
You also need to consider the following:
#
Since Z-units do not interfere with hybridization specificity, all general guidelines for designing primers and probes apply.
Remember that ZNA-units cannot convert poor oligo design into a well performing oligo! ZNA® will improve a given sequence by increasing its affinity to the complementary target sequence mainly due to faster hybridization to the target.
metabion offers ZNA® building blocks of 2 - 5 consecutive ZNA-units spermine units for primers and of 2 – 4 spermine units for probes.
The increase in Tm induced by the introduction of spermine moieties is independent both from the oligonucleotide sequence and from the position of the cationic units (5’ or 3’).
To check your sequence, you can read our FAQ "Are there guidelines that should be taken into account when designing oligonucleotides?".
#
When designing LNA-containing oligonucleotides, one should follow the basic rules of primer design particularly pay attention to the location and number of LNAs.
For example, a typical 18-mer should contain a maximum of 7–8 LNAs.
Also, try to avoid stretches of more than 4 consecutive LNAs, which would result in very tight hybridization in that region.
Stretches of LNAs are to be avoided close to the 3′ end of an oligonucleotide. LNA will bind very tightly to other LNA residues. Avoid self-complementarity and cross-hybridization to other LNA-containing oligonucleotides.
Finally, be sure to match the Tm of the primers, as usual, keeping in mind that each substitution of a standard nucleotide with an LNA increases the Tm by 2-6 °C per LNA.
For novel applications, design guidelines may have to be established empirically by creating your own “best recipe”.
#
Yes, we do! Please, visit our dedicated page.
To learn more about CRISPR-Cas, click here.
#
You may want to consider the following:
#
For a detail explanation of DLPs and their use in qPCR, please refer to our Knowledge Hub.
You also need to consider the following:
#
In principle, antisense oligonucleotides can be synthesized as unmodified phosphodiester oligos. However, these are likely to be rapidly degraded in cells by nucleases. Therefore, antisense oligonucleotides are usually synthesized as chimeric oligonucleotides. metabion offers:
- Phosphorothioate (PTOs)/phosphodiester chimera, which typically have a central core of unmodified DNA and one to four S-modified internucleoside linkages at both 5’ and 3’ end. At least 3 PTO bonds at the 5’ and at the 3’-end of an oligonucleotide are generally recommended to prevent degradation by exonucleases. Adding PTO linkages in the central core of your oligonucleotide (thus making a full-PTO oligonucleotide) will additionally prevent endonucleases-related degradation. However, this might also lead to cytotoxicity effects;
- DNA/ 2'-O-Methyl RNA bases;
- replacement of dC with 5-Methyl-dC, especially in the context of a CpG motif.
Phosphodiester and phosphorothioate oligos are made using a DNA synthesizer, which is basically a computer-controlled reagent delivery system. The first base is attached to a solid support, usually a glass or polystyrene bead, which is designed to anchor the growing DNA chain in the reaction column. DNA synthesis consists of a series of chemical reactions.
Each cycle of reactions results in the addition of a single DNA base. A chain of DNA bases can be built by repeating the synthesis cycles until the desired length is achieved.
#
metabion produces nGS oligos in a dedicated synthesis pipeline. This dramatically minimizes the risk of cross-contamination and guarantees an unmatched degree of high quality. With our nGS oligos, you can perform your Next Generation sequencing experiments with the peace of mind that you work with reagents you can trust.
Oligos are made using a DNA synthesizer which is basically a computer-controlled reagent delivery system. The first base is attached to a solid support, usually a glass or polystyrene bead, which is designed to anchor the growing DNA chain in the reaction column. DNA synthesis consists of a series of chemical reactions.
Each cycle of reactions results in the addition of a single DNA base. A chain of DNA bases can be built by repeating the synthesis cycles until the desired length is achieved.
#
Click chemistry can be used to label both, DNA and RNA oligos of your choice. The synthesis chemistry follows that of standard DNA/RNA oligos. For more information, please refer to our FAQs.
Additionally, an alkyne site is introduced internally or at the 5’/3’ end in the following fashion:
- via an Alxyl moiety, which is sequence independent and allows the addition of an Alkyne group via a C6 (hexynyl)-linker to the phosphate of the 5´ nucleoside;
- via an Alrol moiety, which is sequence independent and allows the addition of an Alkyne group via a serinol-linker to the 3’ OH of the 3´ nucleoside;
- via replacing a C or T at any position of your DNA or RNA oligonucleotide (internally or at the 5’/3’ end) with AldC/AldU, respectively (sequence dependent). AldU and AldC carry a C8-Alkyne motif at the C5 position of the respective Pyrimidine structure of either dU (replacing dT) or dC.
“Ready-to-use/fully labeled” custom oligonucleotides are coupled to the Azide modifier of your choice (please refer to our product portfolio for a full list of our Azide modifiers to use in the “click reaction” described by Carell and colleagues.
#
In accordance with the FRET concept, our DQPs are optimized for real-time quantitative PCR assays, while addressing the need to efficiently mitigate the background noise inherent in probes longer than 25 nts.
Two selection criteria are essential for the length of a linear hydrolysis probe used in 5'-nuclease assays: quenching efficiency and binding stability. As such, sequence design is a careful trade-off between these two factors.
In some cases, to achieve the necessary and acceptable probe Tm for best assay performance, it is necessary to extend the probe sequence beyond the favorable 20-30 nt window, i.e. for AT-rich target sequences. However, as the sequence length increases, the quenching efficiency decreases, leading to enhanced baseline fluorescence and reduced signal-to-noise values for the designed assays.
To overcome this upper probe length limit of 30 nts, it is possible to introduce an internal quencher closer to the fluorescent 5´moiety, while the additional 3'-quencher not only enhances quenching efficiency but also serves as a polymerase inhibitor, thereby avoiding probe extension during hybridization.
Typically, a DQP's internal quencher is located between the 9th and 10th nt downstream of the 5'-fluorescent reporter. This minimizes the distance between the reporter and the quencher and thus the background fluorescence.
R: Reporter/Fluorescent Dye; NFQ: Non-Fluorescent Quencher
#
Please have a look at the following table:
#
A concentration of 0.2 µM of the DQP should be fine for most assays, but as usual the optimal concentration should be determined through your respective experimental setup.
#
Yes, this is possible. Please mark the respective phosphorothioate bond by an asterix (*).
A fully thioated LNA 5´ATCGAT3´ Hexamer for example shall be depicted as follows:
5´ (+A) *(+T) *(+C) *(+G) *(+A) *(+T) 3´
#
A concentration of 0.2 µM of the LNA probe should be fine for most assays, but as usual the optimal concentration should be determined experimentally. Due to the higher affinity of the probe to the target, the probe concentration might be lower.
#
A concentration of 0.2 µM of the MGB probe should be fine for most assays, but as usual the optimal concentration should be determined experimentally. Due to the higher affinity of the probe to the target, the probe concentration might be lower.
#
Oligonucleotides are at the heart of some of the most powerful molecular biology techniques like PCR, DNA chips and in situ hybridization. Moreover, lots of hope has been pinned for oligonucleotides as a generic, yet very selective, class of drugs if they were able to cross cell membranes. Many chemical modifications have been developed, including thiophosphate, peptide nucleic acid, locked nucleic acid and morpholino oligonucleotides, to improve their properties. Owing to the polyanionic nature of oligonucleotides/nucleic acids, conjugation to a polycationic moiety is the rational approach to decrease charge repulsion and thus improve hybridization properties. Moreover, polycations are known to carry oligonucleotides into cells, hence their conjugates are expected to behave similarly. Various synthetic approaches for introducing cations into oligonucleotides have been described, including phosphate backbone replacement, nucleotide modifications or end-conjugation, for which enhanced hybridization, strand invasion and eventually cell penetration were observed.
Among cation-modified oligonucleotides, those leaving the oligonucleotide moiety intact retain mismatch discrimination and remain substrates of nucleic acid–processing enzymes. Moreover, molecular biology applications require rapid on-demand synthesis of oligonucleotide sequences, and this requirement was not fulfilled by previous synthetic approaches.
By conjugation of spermine derivatives as cationic moieties (Z-units) to – by nature – polyanionic oligonucleotides, a versatile concept to increase affinity of oligonucleotides for their target by decreasing electrostatic repulsions has been developed and proven.
ZNA oligos result from the conjugation of a polycationic moiety to an oligonucleotide. This decreases the charge repulsion between the oligonucleotide and its target sequence, both naturally anionic.
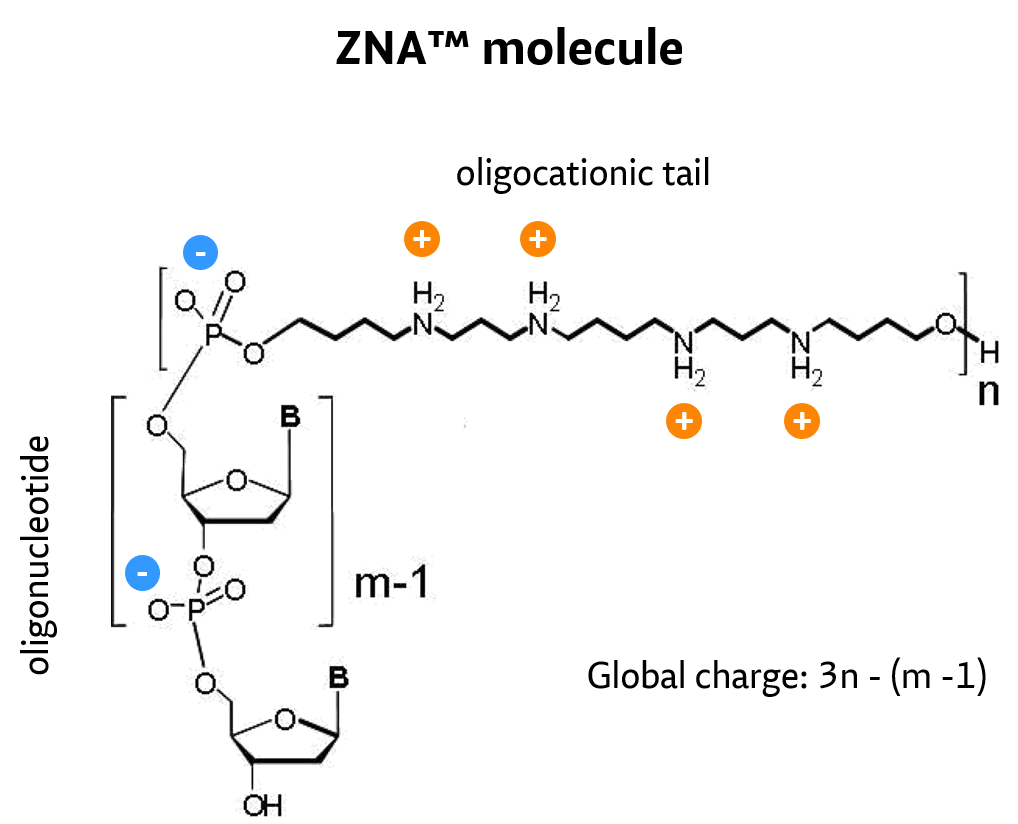
#
ZNA®-synthesis follows standard solid phase, machine-compatible phosphoramidite chemistry. Spermine units are chemically introduced during oligonucleotide synthesis using an appropriately protected spermine phosphoramidite.
#
ZNA®s are delivered in yield ranges for increased transparency and easy calculation of the quantity needed and to be expected. Final yield range is the actual amount that we guarantee to deliver. For example, for a yield range of 5–10 nmol, we guarantee to deliver the final product in a yield between 5 and 10 nmols. See the table below:
*based on oligonucleotides of 8-40 bases in length.
Please note that OD260 values are a measure of total nucleotides´ optical density. Hence, neither purity nor amount of ordered substance are transparently reflected. For simplification and exemplification reasons look at the following:
1 OD of the 20mer 5´CAT CGT ATT CGA TGC TAC GT 3´
translates into approximately 5 nmol.
1 OD of the 40mer 5´CAT CGT ATT CGA TGC TAC GT CAT CGT ATT CGA TGC TAC GT 3´
translates into approximately 2.5 nmol.
Therefore, a 1 OD guaranteed amount of delivered product can vary significantly, while metabion´s commitment to delivered yields in nmol does not allow for ambiguity in terms of what you expect and pay for.
#
The relative Tm increase induced by ZNA or other duplex stabilizing groups is naturally larger on shorter oligonucleotides used as primers or probes. Hence, we offer ZNA primers as short as 10mers and dual-labelled probes as short as 8mers!
The Tm increases significantly and quite linearly with the number of grafted ZNA® spermine units. The approximate Tm of a ZNA® can be calculated applying the following equation:
Tm (ZNA®) = Tm (DNA) + 36z/(N-3.2) (1)
z: number of cationic units
N: number of nucleotides
Example:
Sequence 5´ATATATAT 3´ 8mer Tm(DNA) = 16 °C
plus
ZNA-2 building block
Tm(ZNA) = 16 + 36*2/(8-3.2) = 31 °C
ZNA-2 modification almost doubles Tm; Tm increase of approx 15°C!
Paying respect to the global charge of the ZNA-oligonucleotide-oligocation conjugates raising solubility issues, we additionally offer ZNA-2 and ZNA-3 (cationic) building blocks for (anionic) primers ranging from 8 to 15mers, and dual labeled probes ranging from 10 to 17mers, respectively. Attachment of ZNA-4 and ZNA-5 building blocks to primers is allowed from 16mers (ZNA-4), and 20mers (ZNA-5), respectively. Attachment of ZNA-4 and ZNA-5 building blocks to dual labeled probes is allowed from 18mers (ZNA-4), and 22mers (ZNA-5), respectively.
Considering the ZNA solubility issues and the global charge of a ZNA-oligonucleotide conjugate, a ratio of 1 spermine each 4 nucleotides is appropriate. Accordingly, we offer ZNA-4 building blocks for an unmodified 16mer, for example. In case of dual labeled probes, ZNA-4 is most suitable for a 18mer.
The Tm can be additionally increased, while maintaining specificity, by incorporation of our base analogues:
- C-5 propynyl-dC (pdC) raising melting temperature by ~2.8 °C per substitution
- C-5 propynyl-dU (pdU) raising melting temperature by ~1.7 °C per substitution.
Please note:
There are many different algorithms to calculate the Tm of an oligo. All are just approximations of the actual Tm of a specific oligo under specific conditions (salt concentrations, pH, temperature, sequence composition, oligo length and other biophysical/ biochemical parameters and reaction conditions). Optimization is always recommended.
References
#
ZNA®s display high sensitivity in PCR applications, due to enhanced kinetics in target recognition and “scanning”. This is the reason why ZNA® primers and probes should be used in lower concentration, compared to other DNA-oligos. Typical concentrations are 100-200 nM for primers and 200 nM for probes. A concentration even lower than 50 nM have been proven successful in PCR, without lost of sensitivity. This may represent a great advantage in multiplex assays, which use multiple sets of primers.
#
Due to their lower anionic nature (lower negative charge), compared to pure nucleic acids, ZNA® may be less soluble than other DNA/RNA-oligos in PCR grade water. The solubility of ZNA®s depends mainly on the ratio between nucleotides (anionic)/ spermines (cationic), pH and salt concentration. To avoid solubility issues, ZNA® oligos are delivered already dissolved by default..
If during your experiments you have to dry and re-dissolve your ZNA®, we recommend TE buffer pH 7.4. In case of solubility problems, we recommend adding 50 µl of 50 mM NH4 OH stepwise. Normally, the first 50 µl aliquot is already sufficient to bring a ZNA® oligo into solution. Standard stock concentration for ZNA®s is usually the same as for other PCR primers (100 µM is the delivered concentration). Working solutions of 10 µM should only be used for a short time and ideally prepared instantaneously prior to application.
#
All ZNA® oligos will be delivered at a set concentration of 100 µM, dissolved in H2O (pH 7–9). For special requests, please inquire.
#
For desalted RNAs falling into the specifications of our standard portfolio, you can expect at least >80% of purity. HPLC-purified RNAs falling within the specifications of our standard portfolio, have typical purity >85%. Please note that most delivered RNAs will have purities >90%.
#
Desalted
4 yield ranges based on length-independent delivered quantities from 10-100 nmol for single-stranded oligos from 5-40 nucleotides.
- ≥ 10 < 25 nmol
- ≥ 25 < 50 nmol
- ≥ 50 < 100 nmol
- ≥ 100 nmol
HPLC-purified
5 yield ranges based on length independent delivered quantities* from 5-400 nmol for oligos from 2-80 nucleotides:
- ≥ 5 < 25 nmol
- ≥ 25 < 50 nmol
- ≥ 50 < 100 nmol
- ≥ 100 < 200 nmol
- ≥ 200 < 400 nmol
* Please note that the term “yield range” refers to the final amount of product you will actually receive. Instead, OD260 values are a measure of total nucleotide optical density. Hence, neither purity nor amount of ordered substance are transparently reflected. For simplification and exemplification reasons look at the following:
1 OD of the 20mer 5´cau cgu auu cga ugc uac gu 3´ translates into approximately 5 nmol;
1 OD of the 40mer 5´cau cgu auu cga ugc uac guc auc gua uuc gau gcu acg u 3´ translates into approximately 2.5 nmol.
Therefore, a 1 OD guaranteed amount of delivered product can vary significantly, while metabion´s commitment to delivered yields in nmol for RNAs ranging from 2 to 80 nucleotides does not allow for ambiguity in terms of what you expect and pay for, not even for modified oligos!
#
Distilled sterile water pretreated with DEPC, or any nuclease free biological buffer (i.e. with physiological pH) are acceptable as diluents. The recommended diluent volume is 100 µl - 1 ml, the concentration depending on the application to be used and the yield of the resulting product (see also FAQ "How stable is my RNA DLP once I have resuspended it?").
Please see also our RNA-handling guidelines!
#
Purified sterile water, TE or any biological buffers (i.e. with physiological pH) are acceptable as diluents. The recommended diluent volume is 100 µl - 1 ml, the concentration depending on the application to be used and the yield of the resulting product. For application in cells, we do recommend to use sterile water/buffer. The use of a sterile diluent will also increase the stability of your antisense/PTO. For more information, please read FAQ "How stable is my antisense/PTO once I have resuspended it?".
#
RNA duplexes are significantly more resistant to nucleases than a single strand RNA oligonucleotide. However, maintaining sterile, RNAse free conditions is always recommended as a precaution. Dried pellets are stable at room temperature for 2–4 weeks, but should be placed at –20 °C or –80 °C for long term storage. Under these conditions, dry RNA is stable for at least one year. If you want to store your RNA in solution re-suspend the delivered pellet in an RNAse-free solution buffered at pH 7.4 – 7.6 and store at –20 °C or less. We recommend that RNAs are re-suspended at a convenient stock concentration and stored in small aliquots to avoid multiple freeze thaw cycles.
#
To gain a maximum shelf life for oligonucleotides, samples
should generally be stored dehydrated at ≤ –15 °C in absence of light. Under the
mentioned conditions, samples are stable for at least 6 months. In case of a
longer storage period, oligos should be pretested for molecular integrity prior
to experimental use. If
stored frozen at –20 °C or –70 °C, it will remain stable for several months.
Repeated freeze-thaw cycles should be avoided, as this will denature the dsDNA.
Moreover, the oligo stability in solution depends on the pH. Dissolving an
oligo into acidic solutions may result in its degradation. Therefore, avoid the
use of non-sterile distilled water as a diluent, since solution pH may be as
low as 4–5.
In addition to what above advised, we recommend that you minimize the exposure
of modified antisense/PTOs– especially if fluorescently labelled - to light, to
avoid any bleaching effect.
Moreover, we recommend storing dye-labelled antisense/PTOs highly concentrated and not in working dilutions, if you are not planning to use it within 24 hours. The higher the dilution factor, the faster the fluorescent activity fades away. Therefore, try to store highly concentrated aliquots frozen, thaw them only once, dilute them just before you use the probe and store the aliquots at 4 °C in the dark.
#
Our default delivery form is dry. This ensures maximum stability during transport and storage.
#
Shipping condition dry or wet in 6 different normalization options free of charge:
- Standard desalted, “ready-to-use” and 100% QCed oligos at top quality.
- Minimum number of oligos of the same kind (scale, modification, normalization, shipping condition) is 24 per 96well plate. Combination of batches of >/=24 congeneric oligos in one plate is allowed (e.g. 4x24, 2x48, 3x24, 2x24/1x48 etc.).
#
RNA duplexes are significantly more resistant to nucleases than a single strand RNA oligonucleotide. However, maintaining sterile, RNAse free conditions is always recommended as a precaution. Dried pellets are stable at room temperature for 2–4 weeks, but should be placed at –20 °C or –80 °C for long term storage. Under these conditions, dry RNA is stable for at least one year.
#
Yes, we do! This is part of our standard RNA portfolio! Delivery/shipping condition of dsRNA like for all our RNA oligos is “dry”. Simply dissolve the pellet in annealing buffer and apply a suitable annealing protocol. For your convenience we recommend to follow our guidelines on our download section.
#
All oligonucleotides, whether single-stranded or double-stranded oligos, are provided as dried pellets and shipped at ambient temperature. While being stable at room temperature for 2–4 weeks, they should be placed at –20 °C or at lower temperature upon receipt.
#
The current maximum length of standard RNA Longmers is 140 bases. For RNA Longmers longer than 140 bases, please inquire.
#
RNA duplexes are significantly more resistant to nucleases than a single strand RNA oligonucleotide. However, maintaining sterile, RNAse free conditions is always recommended as a precaution. Dried pellets are stable at room temperature for 2–4 weeks, but should be placed at –20 °C or –80 °C for long term storage. Under these conditions, dry RNA is stable for at least one year.
#
2 ranges of quantity delivered based on length independent delivered quantities* from 10–50 nmol for oligos from 5–40 nucleotides:
- ≥ 10 < 25 nmol
- ≥ 25 < 50 nmol
* Please note that the term “yield range” refers to the final amount of product you will actually receive. Instead, OD260 values are a measure of total nucleotides' optical density. Hence, neither purity nor amount of ordered substance are transparently reflected. For simplification and exemplification reasons look at the following:
1 OD of the DNA 20mer 5´cau cgu auu cga ugc uac gu 3´ translates into approximately 5 nmol;
1 OD of the DNA 40mer 5´cau cgu auu cga ugc uac guc auc gua uuc gau gcu acg u 3´ translates into approximately 2.5 nmol.
Therefore, a 1 OD guaranteed amount of delivered product can vary significantly, while metabion's commitment to delivered oligonucleotides in nmol does not allow for ambiguity in terms of what you expect and pay for, not even for modified oligos!
#
Using our optimized production pipeline, we can deliver RNA DLPs of over 40 bases. The maximum length of an oligonucleotide depends both on the sequence and on the modifications of interest. For example, consider a DLP having a reporter at the 5’-end and a quencher at the 3’-end. The length of the DLP can compromise the activity of the quencher, if the distance between reporter and quencher is too high. Therefore, we consider a length of 40 bases as maximum, for a DLP probe to function optimally.
To get best performance for your experiments, we can help you designing your probe. Please do not hesitate to contact us!
#
Maintaining sterile, RNAse free conditions is always recommended as a precaution. Dried pellets are stable at room temperature for 2–4 weeks, but should be placed at –20 °C or –80 °C for long term storage. Under these conditions, dry RNA is stable for at least one year.
For correct storing and best performance of your RNA DLP, we recommend the following:
- Avoid repeated freeze-thaw, as this will denature the RNA DLP;
- Avoid the use of water as a diluent, since its pH may be as low as 4–5;
The RNA DLP stability in solution depends on the pH. Dissolving DLPs into acidic solutions may result in oligo degradation. Therefore, avoid the use of non-sterile distilled water as solution pH might be as low as 4–5; - Minimize the exposure of RNA DLPs to light, to avoid any bleaching effect;
- Store RNA DLPs highly concentrated and not in working dilutions, if you are not planning to use them within 24 hours. The higher the dilution factor, the faster the fluorescent activity fades away. Therefore, try to store highly concentrated aliquots frozen, thaw them only once, dilute them just before you use the probe and store the aliquots at 4 °C in the dark;
- For general instructions on how to handle RNA, please refer to our Guidelines for handling RNA.
#
Just to avoid any potential misconception concerning the production technique applied, building dsDNA by adding complementary base pairs step by step is not possible. The method of choice consists in synthesizing two single strand DNA (ssDNA) oligos separately and align them to form a duplex via complementary base pairing. This drives the formation of more or less stable duplexes/dsDNA.
Each ssDNA oligo is made using a DNA synthesizer, which is basically a computer-controlled reagent delivery system. The first base is attached to a solid support, usually a glass or polystyrene bead, which is designed to anchor the growing DNA chain in the reaction column. DNA synthesis consists of a series of chemical reactions.
Each cycle of reactions results in the addition of a single DNA base. A chain of DNA bases can be built by repeating the synthesis cycles until the desired length is achieved.
#
The real answers lies in the resolution limit of the purification method and on the coupling efficiency of the DNA synthesizer. We can synthesize dsDNA/duplex DNA of 220 bases and obtain sufficient quantities by HPLC purification to perform successful gene construction. However, it should be remembered that the longer the DNA construct, the greater the chance of accumulated sequence errors.
Coupling efficiency is the factor that mainly affects the length of DNA that can be synthesized. Base composition and synthesis scales will also be contributing factors. The table below shows that at 99% coupling efficiency, a crude solution of synthesized 95-mers would contain 38% full-length product and 62% (n–x) failure sequences. This is without taking into account other chemical reactions, such as depurination (which mainly affects the base A). The frequency of depurination is small but increases significantly with the primer length. However, as we are very experienced in producing very long oligos, do not hesitate to discuss your projects with our specialists.
#
Each siRNA molecule is synthesized as two ssRNA oligonucleotides between 11 and 40 nucleotide in length, which are then aligned to form a duplex. The siRNA duplex typically has 2 DNA nucleotides overhanging at each 3’-end. For more information about RNA synthesis, read our FAQs.
Each ssRNA strand is HPLC purified as well as mass-checked for best quality.
#
There are two ways of ordering:
- The preferred way is order transmission through our Web Order Portal for most convenient online shopping (please refer here for further details).
- You can order by sending us an e-mail at oligo@metabion.com with our pre-formatted excel order file as attachment. Download respective Order Form.
When you write your e-mail, please make sure to include the following required information in the excel template:
- Name of the siRNA/s.
- Sequence of the siRNA/s in 5’-3’ orientation.
- Yield range.
- Purification required.
- Delivery form (dry).
- Modifications.
If you are a new customer, please additionally provide us with:
- Your shipping and billing address.
- Any other information like Purchase Order number, VAT number (VAT only for customers resident in the EU), etc.
In case you choose to transmit orders via e-mail using your own format(s), we need to alert you that above mentioned information in bold print is obligatory for processing your order. Due to extra efforts necessary for individual order format transfer into our system, order processing will take longer as compared to preferred web orders and pre-formatted e-mails. Please note that only files updated to the latest excel/word version (e.g. .xlsx or .docx) are accepted.
How can you convert files to newer formats?
Open
the file in a recent Office program and save the file in a newer format.
For this go to “File” → “Save as” and choose the newer format from the
“Save as type” dropdown menu.
If you want to connect your eProcurement System with our Web Order
Portal (e.g. OCI - Open Catalog Interface), please simply contact our
Customer Service (customerservice@metabion.com).
#
You may want to consider the following:
#
The expected average in-house turnover time for siHE siRNAs is 6–8 working days. Please note that we perform strict quality controls on each and every oligo. In case one or more oligos do not pass our quality control, they will need to be resynthesized. This may, of course, result in a delay.
In terms of "counting" working days, orders placed past 15:00 (Munich time) are considered to be next day's order.
#
siHE siRNAs are provided as dried pellets and shipped at ambient temperature. While being stable at room temperature for 2–4 weeks, they should be placed at –20 °C or at lower temperature upon receipt.
#
The label on the siHE siRNA tube shows basic information like oligo name, name of person who placed the order, siRNA sequence including modifications, oligo ID, amount of RNA (OD260 and nmol), Tm, and molecular weight.
In addition, you will receive a technical data sheet containing more detailed information on the physical-chemical properties of the oligo, such as base composition, base count, purification grade, amount of RNA in nmol, Tm and molecular weight. Additionally, you will also receive a hard copy of the Mass-Check documentation/spectrum of each single strand. The following terminology is used for differentiating between offered QC options including respective documentation coverage in our order forms and on supporting documents delivered with the products:
Mass Check
Standard quality control performed on each and every oligo. Either MALDI- or
ESI-ToF, subject to the "nature of the oligo", and metabion internal
procedures. This service is free of charge and the Mass Check spectrum will be
provided.
MALDI-ToF
Explicitly ordered and performed MALDI-ToF check. Product delivered with
MALDI-ToF spectra. Additional charges apply.
ESI-ToF
Explicitly ordered and performed ESI-ToF check. Product delivered with ESI-ToF spectra. Additional charges apply.
#
- Enhanced efficiency in gene silencing compared to standard siRNAs:
- Meticulously designed and synthesized to target specific genes with unparalleled accuracy;
- Ensures precise and highly efficient gene silencing;
- Allows researchers to dissect complex biological pathways with ease.
- Custom options to meet specific research needs.
- Rigorous quality control measures ensure purity and efficacy.

